Understanding the Mode of Action of a Compound
To help clients understand the mode of action of their compounds, we use tools such as gene expression analysis, gene editing and custom (bespoke) assays.
Gene Expression Analysis
When a compound’s mechanism of action is unknown, we can clarify it by examining differential gene expression, either on a targeted scale using qRT-PCR or on an omics scale using RNA-seq. We have the computational expertise and tools to develop biologically meaningful hypotheses from differential gene expression signatures for RNA-seq.
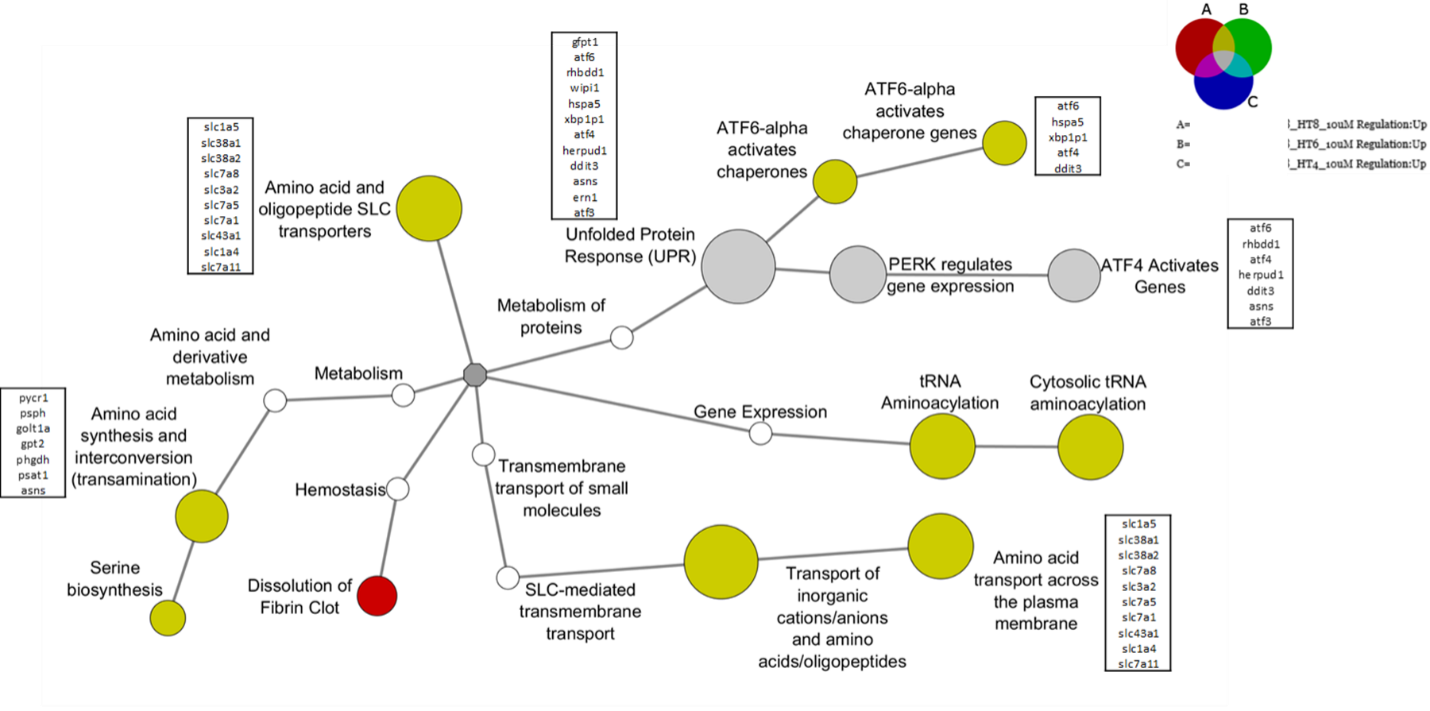
Differential gene expression analysis: Drug treatment caused up-regulation of genes involved in the indicated biological processes. Color indicates which experimental conditions induced upregulation (red = 8 h treatment, yellow = 6 h and 8 h treatments, gray = 4 h, 6 h, and 8 h treatments).
Hypothesis Testing with RNAi and Gene Editing
Transcriptomics is an excellent technique for hypothesis development, however, testing hypotheses frequently relies on manipulations of protein expression levels through overexpression, knockdown, or knockout. Our experience lies in using lentiviral transduction to generate stable cells lines overexpressing wild-type or mutant proteins, expressing shRNAs for stable knockdown of target proteins. We also use the CRISPR/Cas9 system to generate cell lines with homozygous knockout of target proteins. These cell lines allow us to test the roles of proteins and pathways of interest in pharmaceutical function.
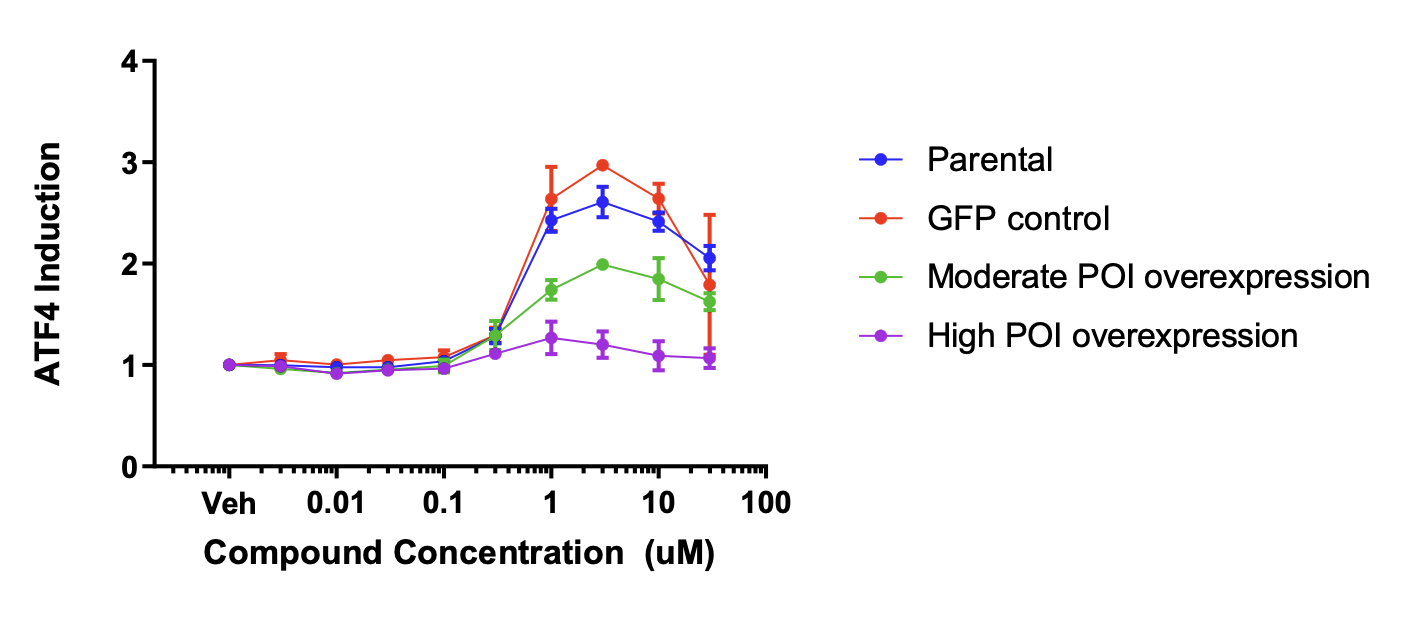
The figure above shows drug activity is modulated by expression level of a protein of interest.
Custom Assays
To investigate a specific mode of action for a compound, we leverage our knowledge of the biology to design cell-based experiments to measure the relevant endpoint. For example, a client was concerned that their chemistries were causing liver toxicity through fatty acid build-up (a phenomenon) known as steatosis. We developed an assay to specifically measure the fatty acids that are chemically induced. The uterine assay is another example of a bespoke assay that we developed.

Case Study: Using Transcriptomics To Determine Interspecies Differences In The Mode Of Action
Case Study: Understanding The Mode Of Action Of Lead Candidates
Case Study: Understanding The Mode Of Action Of A Food Contaminant Acetamide
